The Problem
• It is estimated that 1.8 billion people are systematically exposed to contaminated drinking water sources.
• Water quality can deteriorate at any stage of its course, from water springs to inadequate treatment strategies and deteriorated piping.
• The presence of infectious agents at high microbial loads is the main cause of disease in these cases, with Nepal being a dire example where waterborne diseases account for 15% of all health cases.
• Inadequate urban wastewater treatment can exacerbate the problem by adding harmful microorganisms to aquifers.
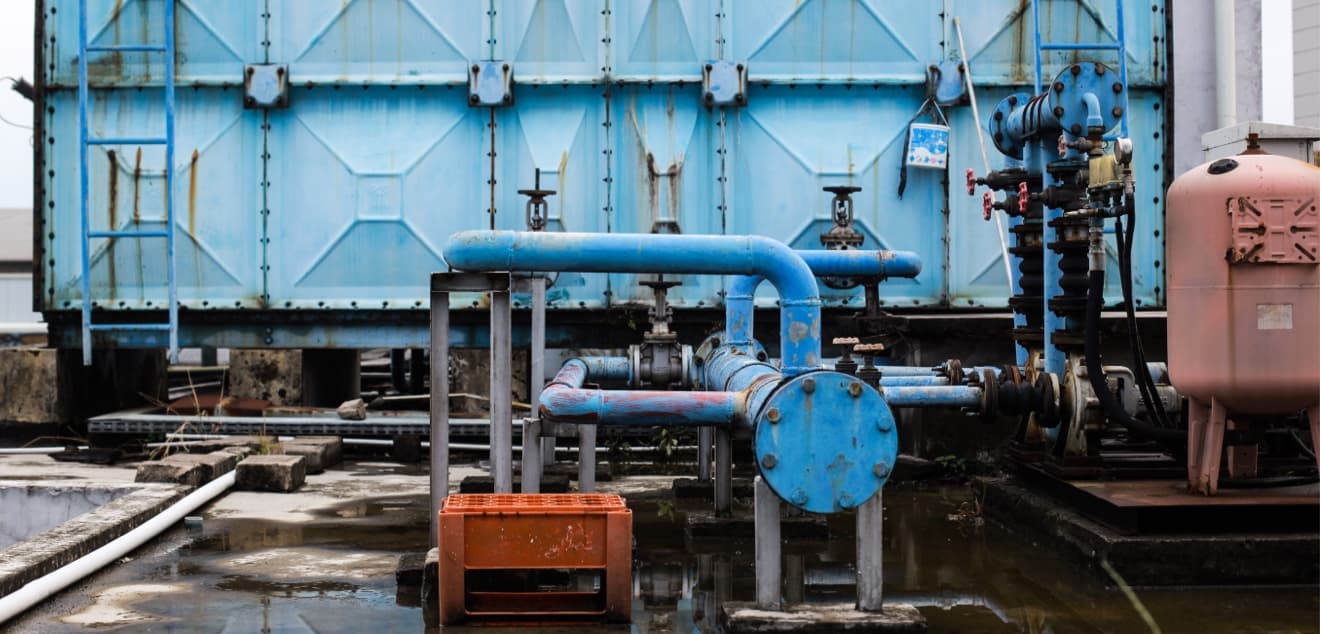

Proposed solution
Our main goal is to detect and systematically monitor microbial communities found in closed water systems and wastewater with the ultimate goal of ensuring water quality and safety.
By constructing almost complete genomes from metagenomic samples, we are able to exhaustively identify all potential biological hazards and monitor their evolution and spread during periods of epidemic outbreaks (eg SARS-CoV2).
Monitoring pathogens in the environment in almost real time allows targeted and effective infection control measures.
Monitoring the microbial load in closed water systems and in wastewater will lead to the development of better strategies for pathogen spread mitigation.
Monitoring

Drinking water

Water supply system

Urban sewage

Effluent
Benefits of the analysis

Identifying and monitoring the microbial footprint of urban wastewater and sewage is an indicator of epidemiological burden and can be used by the competent authorities both for the protection of public health. (as in the case of SARS-CoV2) and for sanitary waste management.

Microbial footprint identification in water supply systems can be a key indicator of the quality of drinking water.

The MBE solution of DNA sequence provides quasi real time information on possible water supply systems health threats, so that the necessary measures can be taken in a timely manner.

The interactive result reports generated by the DNA sequence Biosafety platform can be used to inform authorities and interested third parties.

